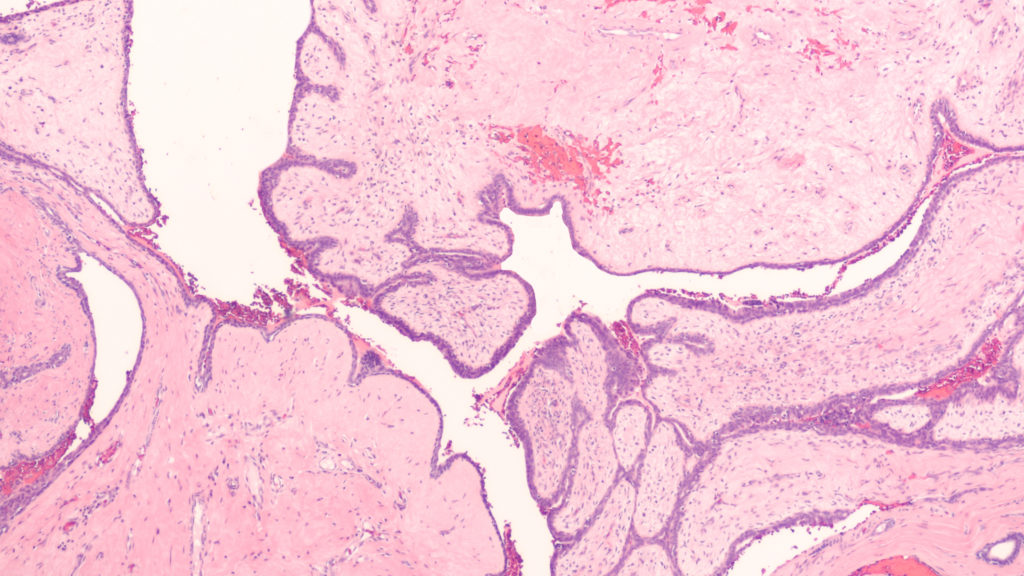
Researchers conducted a 2021 study to better characterize phyllodes tumors and other breast fibroepithelial lesions in order to improve diagnosis and treatment for patients.
The Trending with Impact series highlights Oncotarget publications attracting higher visibility among readers around the world online, in the news, and on social media—beyond normal readership levels. Look for future science news and articles about the latest trending publications here, and at Oncotarget.com.
—
Thankfully, around 80% of lumps found in human breasts turn out to be benign, or indolent, fibroadenoma (FAD). FADs fall into a category of breast fibroepithelial lesions (FELs), which include many heterogeneous pathological tumors, ranging from benign FADs to rare and potentially aggressive phyllodes tumors (PTs). After examination by a physician, these FELs may be diagnosed as either benign, borderline, or malignant.
However, there is a need to improve accurate diagnosis and distinction between FELs using a marker-based diagnostic approach. In an effort to better characterize FELs, researchers from India’s CSIR-Centre for Cellular and Molecular Biology, Institute of Bioinformatics, Gandhi Hospital, Government Medical College, and Manipal Academy of Higher Education conducted a trending 2021 study, titled: “Quantitative proteome profiling stratifies fibroepithelial lesions of the breast.”
“The current grading system remains unreliable in differentiating these tumors due to histological heterogeneity and lack of appropriate markers to monitor the sudden and unpredictable malignant transformation of PTs.”
The Study
To begin identifying the differentially expressed genes and proteins among FADs and PTs in benign, borderline, and malignant states, the researchers conducted quantitative global proteomics on Formalin-Fixed Paraffin-Embedded (FFPE) tissue sections. They conducted a principal component analysis of the protein expression matrix to identify the overlapping proteomic profiles among FELs.
“Interestingly, we observed FADs and benign PTs clustered together compared to borderline and malignant ones, albeit with overlapping protein expression profiles.”
When FADs were compared with benign PTs, the researchers identified 32 proteins in FAD that were differentially regulated. The researchers elucidated many important distinctions between benign, borderline, and malignant FADs and PTs, and identified at least three potential prognostic markers that may aid in patient diagnosis and treatment. The progression of PTs from borderline to malignant and their mechanistic framework was clearly explained by the researchers in this study.
“The presence of extensive ECM proteins and EMT markers led us to hypothesize a model of deposition and degradation of these proteins thus triggering ECM remodeling and EMT acquisition in borderline PTs leading to its malignant state. Enrichment of platelet degranulation factors in malignant PT indicates active angiogenesis during this transformation.”
The Study
“Herein, our initial findings suggest that MUCL1, HTRA1, and VEGFD can be used as potential proteomic markers that could augment existing diagnosis, and help in monitoring the progression of the disease.”
Additional characterization of FELs using different omics platforms was recommended by the researchers to help better understand and manage the dynamics of PTs and malignant breast tumors.
“The present work shed light on a brief mechanistic framework of PTs aggressive nature and present potential biomarkers to differentiate overlapping FELs that would be of practical utility in augmenting existing diagnosis and disease management for this rare tumor.”
Click here to read the full scientific study, published in Oncotarget.
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read—without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues, and other researchers, far and wide.
For media inquiries, please contact media@impactjournals.com.