Researchers developed a new tool aimed at better classifying HPV+ HNSCC patients with good or poor prognosis in an effort to personalize treatment and improve patient outcomes.
—
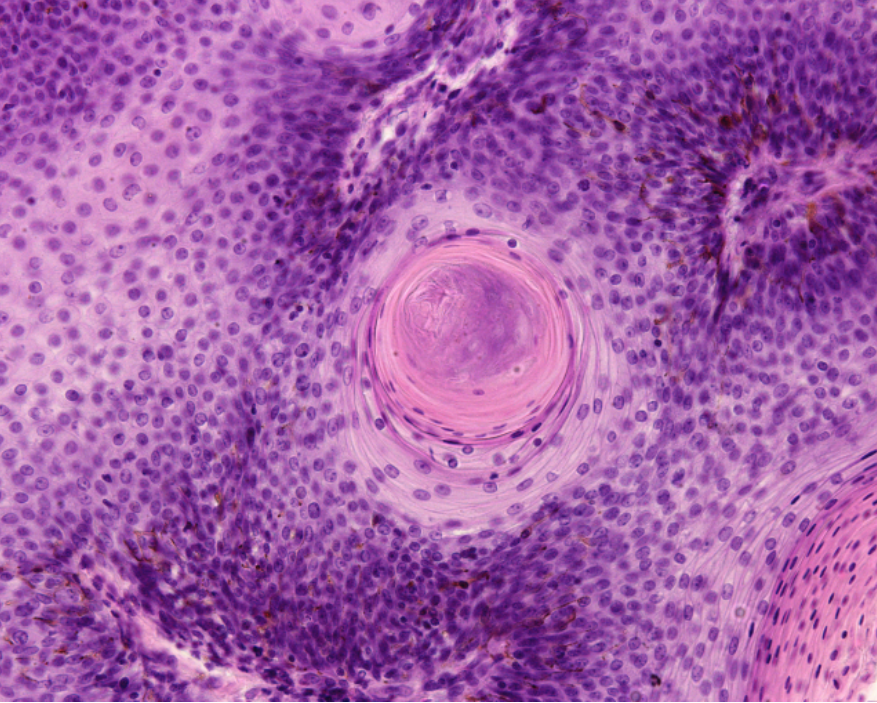
Over the last 10 years in the United States, the human papillomavirus (HPV) has caused more head and neck squamous cell carcinomas (HNSCC) than uterine cervical cancers. Primarily caused either by exposure to HPV or to ethanol or tobacco, HNSCC is a disease that impairs fundamental tissues involved in respiration, speech and digestion. HPV-positive and -negative HNSCC have contrasting clinical, epidemiological and histological features.
“A major discovery in the recent past is that HPV associated HNSCC have improved survival compared to tobacco associated tumors.”
Therefore, treating HNSCC in accordance with HPV status is crucial for avoiding unnecessarily harsh therapeutic side effects in HPV+ HNSCC patients. However, while oncologic outcomes among patients with HPV+ HNSCC are generally favorable, approximately 30% experience a more aggressive disease course and recurrence. Coupled with increasing incidence worldwide, this highlights a growing need for the development of effective clinical stratification tools to accurately identify HPV+ HNSCC patients who have a good or poor prognosis.
In a new study, researchers—from Columbia University, University of Illinois Cancer Center, University of North Carolina at Chapel Hill, and Yale School of Medicine—developed a new tool aimed at better classifying HPV+ HNSCC patients with good or poor prognosis in an effort to personalize treatment and improve patient outcomes. Their trending research paper was published in Oncotarget on May 24, 2022, and entitled, “NF-κB over-activation portends improved outcomes in HPV-associated head and neck cancer.”
“To improve on genomic classification, we designed this study to provide a foundation for development of NF-κB related, RNA based classification strategies to better identify HPV+ HNSCC patients with good or poor prognosis that could potentially aid in future efforts towards treatment personalization.”
The Study
The researchers from this study previously found that TRAF3 and CYLD genes are negative regulators of a family of inducible transcription factors involved in inflammation, called nuclear factor kappa B or NF-κB. The team found that somatic mutations or deletions in either TRAF3 or CYLD (not commonly found in uterine cervical cancer or HPV-negative HNSCC) lead to increased NF-κB pathway activation in HPV+ HNSCC. NF-κB overactivity may lead to cancer cell growth and survival. Alterations in these NF-κB related genes may be potential therapeutic targets in HPV+ HNSCC, and their expression may be capable of predicting treatment outcomes.
“[…] we hypothesized that tumor groups based on NF-κB related gene expression may correlate with treatment outcome, considering that tumors lacking defects in TRAF3 and CYLD may have unrecognized mechanisms driving constitutive NF-κB activation.”
In the current study, the researchers developed an RNA-based NF-κB classification tool called the NF-κB Activity Classifier, or NAC. They used bioinformatics and machine learning techniques, expression-based classification, principal component (PC) analysis, gene set enrichment analysis, and weighted gene correlation network analysis (WGCNA) to verify that the NAC is indeed capable of identifying tumors with high or low NF-κB activity and tumors with good and poor survival.
“This report validates and expands on our findings that significant expression changes related to NF-κB activity occur in the subset of HPV+ HNSCC tumors marked by TRAF3 or CYLD mutations. We are planning future studies investigating the importance of ‘long-tail’ mutations in the NF-κB pathway which might further illuminate the origins of NF-κB dysregulation in HPV+ HNSCC.”
Conclusion
“Here we present data that these subclasses may also be identified by direct assessment of NF-κB activity; as demonstrated by gene expression differences highlighted by the NF-κB Activity Classifier.”
In summary, the researchers identified genomic differences within subclasses of HPV+ HNSCC. They found that defects in TRAF3 and CYLD genes and NF-κB activity were correlated with survival. Therefore, the NF-κB Activity Classifier could be a useful guide for clinicians who make therapeutic decisions for patients with HPV+ HNSCC.
“Future applications of the NF-κB Activity Classifier may be to identify HPV+ HNSCC patients with better or worse survival with implications for treatment strategies.”
Click here to read the full research paper published by Oncotarget.
ONCOTARGET VIDEOS: YouTube | LabTube | Oncotarget.com
—
Oncotarget is an open-access journal that publishes primarily oncology-focused research papers in a continuous publishing format. These papers are available at no cost to readers on Oncotarget.com. Open-access journals have the power to benefit humanity from the inside out by rapidly disseminating information that may be freely shared with researchers, colleagues, family, and friends around the world.
For media inquiries, please contact media@impactjournals.com.