This review compiles splice variations in HER2 breast cancer, specifically in the context of the tumor environment, and co-expression of variants. The study also provides an up-to-date (as of Nov. 2020) account of HER2 and HER2 variant patterns of resistance to anti-HER2 therapies and other interventions.
The Trending with Impact series highlights Oncotarget publications attracting higher visibility among readers around the world online, in the news, and on social media—beyond normal readership levels. Look for future science news and articles about the latest trending publications here, and at Oncotarget.com.
—
According to Cancer Research UK, breast cancer occurs in one in every eight women within their lifetime and is the second highest cause of cancer related deaths in the UK. Breast cancer is a blanketed term for a wide variety of tumors that occur in the mammary glands. In over 20% of breast cancers, the human epidermal growth factor receptor two gene, officially named ErbB2 but otherwise known as HER2, is overexpressed. HER2 is a member of the epidermal growth factor receptor family of EGFR, HER2, HER3, and HER4. Overexpression of the HER2 protein was discovered in 1987 as a biomarker associated with poor prognosis and aggressive tumor types in breast cancer. This finding has accelerated research studies and progress in HER2 diagnostic testing and targeted therapeutics. However, the issue of HER2 resistance in these targeted therapies remains problematic.
“At the present time, we have an incomplete understanding of why patients with HER2+ breast cancer exhibit variable responses or resistance to targeted therapies [73, 74].”
Researchers from the Translational and Clinical Research Institute at Newcastle University in the United Kingdom have compiled a review of variations in HER2 breast cancer, specifically in the context of the tumor environment and when multiple variants are co-expressed at altered ratios. Their study also provides an up-to-date (as of Nov. 2020) account of the current landscape of HER2 variants and links this to patterns of resistance against HER2 therapies and other interventions.
“It is clear HER2 expression is not as simple as a single oncogenic overexpressed protein. It is likely many variants, arising from splicing and other mechanisms, are present in tandem. The relative ratios of these are likely to fluctuate depending on cellular conditions, during tumorigenesis and breast cancer progression.”
HER2 Variants & Co-expression
This paper provides an exquisitely detailed description and explanation of the HER2 protein structure, signaling pathways, sub-typing, and in-depth treatment functionality of a number of different HER2 targeted therapeutics.
“Different forms of the HER2 protein exist within tumours in tandem and can display altered biological activities.”
The unique interest in researching variations in HER2 breast cancer has increased since the identification of Δ16-HER2: a particular splice variant and link to resistance of anti-HER2 therapies. The “Δ16” in Δ16-HER2 refers to the lack of exon-16, which encodes a small extracellular portion of the DNA. Δ16-HER2 represents approximately 9% of the normal HER2 transcripts and its expression is considered common in breast cancer. Previous studies have identified Δ16-HER2 and HER2 normal transcripts can be co-expressed at varying levels in breast carcinomas.
In the variant P100, less is known about this truncated HER2 protein. It has been hypothesized that P100 reduces the efficacy of monoclonal antibody HER2 treatments.
The splice variant Herstatin is produced by the retention of intron-8 in the HER2 protein. Herstatin acts as a tumor suppressor by blocking HER2 activity and cell proliferation, while promoting apoptosis. The researchers mention that it is important to note that cells expressing high levels of Herstatin are more sensitive to Tamoxifen.
“It’s noteworthy that one study proposed that the presence of Herstatin transcript does not segregate by tumour grade or size, patient age, lymph node involvement or ER status and that mRNA transcripts were present in matched non-cancerous breast tissue and breast carcinomas [96].”
Researchers in this review note that identifying and assessing the expression ratios of these different variants and classifying them as prognostic and predictive biomarkers may aid in further personalized treatment of breast cancer in HER2 positive patients.
Testing and Research Landscape
“Studying splicing regulation and how this is altered in breast cancer could explain patterns of expression and how these link to treatment resistance [111].”
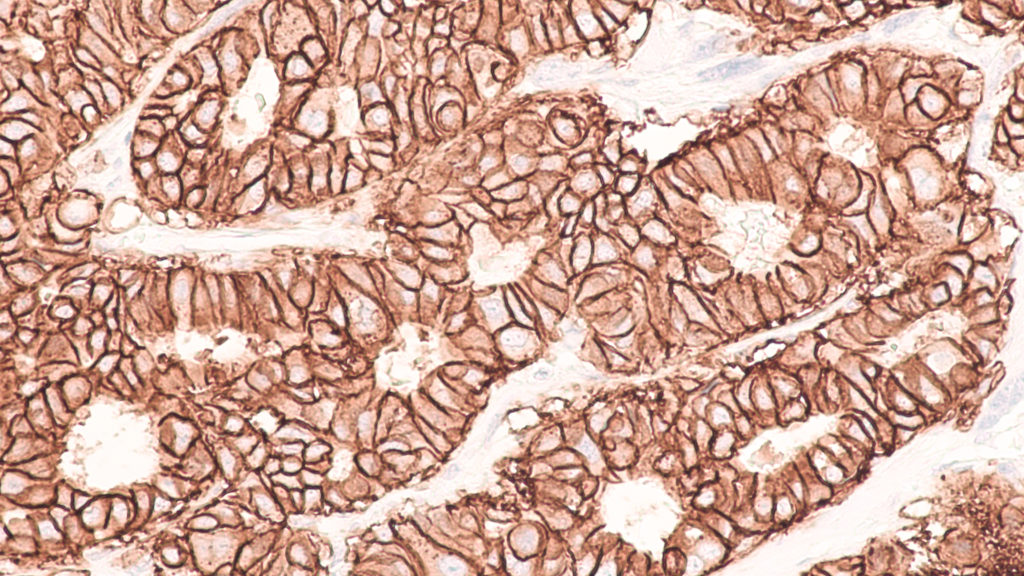
The researchers write that tests assessing for both HER2 status and HER2 variant expression could potentially refine their predictions of a patients’ response to treatment. One common way that researchers gauge levels of HER2 proteins, and only some HER2 variants, is through immunohistochemistry tests. mRNA assessments are also used to identify gene expression patterns. Another biomarker test the researchers noted that may be best used for prognostic predictions is the Enzyme-Immunoassay—to assess levels of plasma or serum HER2 (sHER2) in the blood produced by cleavage or splicing.
“Cohort studies have identified sHER2 testing as a useful complementary test to IHC owing to the correlations between high sHER2 and aggressive tumour phenotypes such as invasion and metastases.”
Targeted HER2 Treatments
The review elaborates in detail about targeted treatments for HER2 breast cancer, which include: trastuzumab, pertuzumab, lapatinib, and T-DM1. They note that endocrine therapy is utilized for ER positive patients and chemotherapy, radiotherapy, and surgery are all still utilized.
Conclusion
“Work in vitro and in vivo as well as analysis from clinical trials has identified patterns of resistance to the standard of care treatment options in HER2+ patients which are correlated to variant expression.”
This goal of their review was to summarise the current landscape of HER2 variant research and to explain why researchers should consider HER2 variant levels and ratios when offering the best treatment plan for breast cancer patients.
Click here to read the full review, published in Oncotarget.
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read—without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues and other researchers, far and wide.
For media inquiries, please contact media@impactjournals.com.
